Essential idea: Meiosis leads to independent assortment of chromosomes and unique composition of alleles in daughter cells.
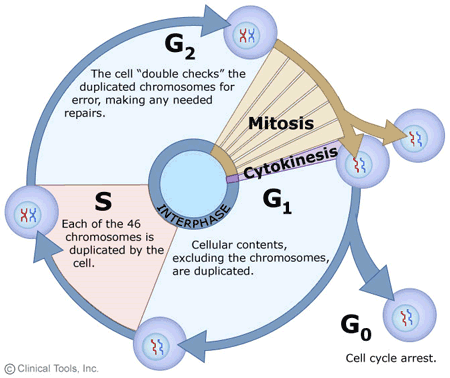
10.1 Meiosis
U10.1.1 Chromosomes replicate in interphase before meiosis.
Chromosomes must replicate in an uncondensed statebefore the cell divides because in order to end up with 4 daughter cells containing 23 chromosomes, the cell must begin with 92 chromosomes, since the cells undergo division twice.
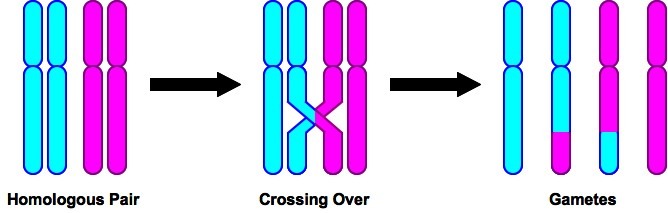
U10.1.2 Crossing over is the exchange of DNA material between non-sister homologous chromatids.
U10.1.3 Crossing over produces new combinations of alleles on the chromosomes of the haploid cells.
U10.1.4 Chiasmata formation between non-sister chromatids can result in an exchange of alleles.
U10.1.1 Chromosomes replicate in interphase before meiosis.
Chromosomes must replicate in an uncondensed statebefore the cell divides because in order to end up with 4 daughter cells containing 23 chromosomes, the cell must begin with 92 chromosomes, since the cells undergo division twice.
U10.1.2 Crossing over is the exchange of DNA material between non-sister homologous chromatids.
U10.1.3 Crossing over produces new combinations of alleles on the chromosomes of the haploid cells.
U10.1.4 Chiasmata formation between non-sister chromatids can result in an exchange of alleles.
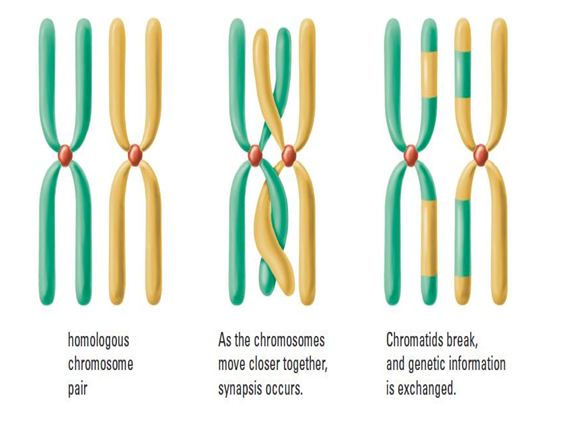
Synapsis is the fusion of chromosome pairs at the start of meiosis. When this happens, crossing over can take place. Crossing over is the exchange of genetic material between non-sister homologous chromatids i.e. the 'matchine' pair of chromosomes.
To sum up:
- Crossing over involves the exchange of segments of DNA between homologous chromosomes during Prophase I of meiosis
- The process of crossing over occurs as follows:
- Homologous chromosomes become connected in a process called synapsis, forming a bivalent (or tetrad)
- Non-sister chromatids break and recombine with their homologous partner, effectively exchanging genetic material (crossing over)
- The non-sister chromatids remain connected in an X-shaped structure and the positions of attachment are called chiasmata
- Chiasma hold homologous chromosomes together as a bivalent until Anaphase I
- As a result of crossing over, chromatids may consist of a combination of DNA derived from both homologues - these are called recombinants (new combinations). The formation of recombinants basically makes the possible allele combinations unlimited.
NOTE: Bivalents are the pairs of the pairs...this is often very misleading - bivalents are NOT sister chromatids - they are the pairs of a pair of homologous sister chromatids i.e. FOUR chromosomes.
U10.1.5 Homologous chromosomes separate in meiosis I.
U10.1.6 Sister chromatids separate in meiosis II.
Prophase I
1. Chromosomes become visible as the DNA becomes more compact i.e. the chromosomes condense.
2. Homologous chromosomes are attracted to (associate with) each other and pair up and become sister chromatids.
3. Crossing over (recomination) takes places between non-sister chromatids - chiasmata is the location at which crossing over takes place on chromosomes.
4. Nuclear membrane disintergrates.
Metaphase I
1. The bivalents (pairs of homologous chromosomes i.e. two sister chromatids) line up at/across the cell's equator.
2. Spindle fibres made from microtubules form.
3. Random orientation takes place.
Anaphase I
1. Spindle fibres from the poles at each end of the cell attach to one sister chromatid and pull them (contract) to opposite poles.
2. Non-disjunction (when pairs of chromosomes do not separate) can occur here and will affect the chromosome number of all four gametes.
Telephase I
1. Spindles and spindle fibres disintergrate.
2. Usually, the chromosomes uncoil and new nuclear membranes form.
Now meiosis II takes place in order to separate the sister chromatids.
Prophase II
1. DNA condenses into visible chromosomes again.
2. No crossing over takes place.
3. Nuclear membranes disintergrate.
Metaphase II
1. Sister chromatids line up at the equator of each cell.
2. Random orientation takes place again.
Anaphase II
1. Spindle fibres form and break the centromeres of each sister chromatid, releasing each as individual chromosomes.
2. Spindle fibres contract and pull chromosomes to opposite poles of the cells.
3. Because of random orientation, the chromosomes can be pulled towards either of the newly forming daughter cells.
4. Non-disjunction can occur here as well.
Telephase II
1. Chromosomes unwind their strands of DNA.
2. Nuclear envelopes form around each of the four haploid cells, preparing them for cytokinesis (cytoplasmic division of each of the two cells).
MEIOSIS = DIPLOID -> DIPLOID -> HAPLOID
U10.1.7 Independent assortment of genes is due to the random orientation of pairs of homologous chromosomes in meiosis I.
APPLICATION:
N/A
SKILL:
S10.1.1 Drawing diagrams to show chiasmata formed by crossing over.
See U10.1.4.
U10.1.6 Sister chromatids separate in meiosis II.
Prophase I
1. Chromosomes become visible as the DNA becomes more compact i.e. the chromosomes condense.
2. Homologous chromosomes are attracted to (associate with) each other and pair up and become sister chromatids.
3. Crossing over (recomination) takes places between non-sister chromatids - chiasmata is the location at which crossing over takes place on chromosomes.
4. Nuclear membrane disintergrates.
Metaphase I
1. The bivalents (pairs of homologous chromosomes i.e. two sister chromatids) line up at/across the cell's equator.
2. Spindle fibres made from microtubules form.
3. Random orientation takes place.
Anaphase I
1. Spindle fibres from the poles at each end of the cell attach to one sister chromatid and pull them (contract) to opposite poles.
2. Non-disjunction (when pairs of chromosomes do not separate) can occur here and will affect the chromosome number of all four gametes.
Telephase I
1. Spindles and spindle fibres disintergrate.
2. Usually, the chromosomes uncoil and new nuclear membranes form.
Now meiosis II takes place in order to separate the sister chromatids.
Prophase II
1. DNA condenses into visible chromosomes again.
2. No crossing over takes place.
3. Nuclear membranes disintergrate.
Metaphase II
1. Sister chromatids line up at the equator of each cell.
2. Random orientation takes place again.
Anaphase II
1. Spindle fibres form and break the centromeres of each sister chromatid, releasing each as individual chromosomes.
2. Spindle fibres contract and pull chromosomes to opposite poles of the cells.
3. Because of random orientation, the chromosomes can be pulled towards either of the newly forming daughter cells.
4. Non-disjunction can occur here as well.
Telephase II
1. Chromosomes unwind their strands of DNA.
2. Nuclear envelopes form around each of the four haploid cells, preparing them for cytokinesis (cytoplasmic division of each of the two cells).
MEIOSIS = DIPLOID -> DIPLOID -> HAPLOID
U10.1.7 Independent assortment of genes is due to the random orientation of pairs of homologous chromosomes in meiosis I.
- Variety produced by recombination of maternal and paternal chromosomes
- Each pair of homologous chromosomes assorts to daughter cells randomly
- Possible arrangements of chromosomes in haploid daughter cells = (2)^nth, where n = number of homologous pairs - in humans n = 23 and possible arrangements = (2)^23 = about 8 million
- Mendel’s law of independent assortment applies only to traits carried on different chromosomes, i.e. unlinked genes
- Independent assortment occurs as a result of the alignment (random orientation) of homologues during Metaphase I, determining which chromosome assort to which daughter cell
- Each pair of alleles separates independently of every other pair of unlinked alleles
APPLICATION:
N/A
SKILL:
S10.1.1 Drawing diagrams to show chiasmata formed by crossing over.
See U10.1.4.
Essential idea: Genes may be linked or unlinked and are inherited accordingly.
10.2 Inheritance
UNDERSTANDINGS:
U10.2.1 Gene loci are said to be linked if on the same chromosome.
Linkage refers to genes that are located on the same chromosome. Linked genes tend to be inherited together and the extent of crossing over depends on how close together they are on the chromosome. Linkage generally reduces the variety of offspring that can be produced. In genetic crosses, linkage is indicated when a greater proportion of the offspring resulting from a cross are of the parental type (than would be expected if the alleles were assorting independently). If the genes in question had been on separate chromosomes, there would have been more genetic variation in the gametes and therefore in the offspring.
U10.2.2 Unlinked genes segregate independently as a result of meiosis.
U10.2.3 Variation can be discrete or continuous.
Discontinuous variation This is where individuals fall into a number of distinct classes or categories, and is based on features that cannot be measured across a complete range. You either have the characteristic or you don't. Blood groups are a good example: you are either one blood group or another - you can't be in between, same goes for gender. Such data is called discrete (or categorical) data. Chi-squared statistical calculations work well in this case.
In continuous variation there is a complete range of measurements from one extreme to the other. Height is an example of continuous variation - individuals can have a complete range of heights, for example, 1.6, 1.61, 1.62, 1.625 etc metres high.
U10.2.4 The phenotypes of polygenic characteristics tend to show continuous variation.
A10.2.3 Polygenic traits such as human height may also be influenced by environmental factors.
Continuous variation is the combined effect of many genes (known as polygenic inheritance) and is often significantly affected by environmental influences. Milk yield in cows, for example, is determined not only by their genetic make-up but is also significantly affected by environmental factors such as pasture quality and diet, weather, and the comfort of their surroundings.
Polygenic traits are controlled by two or more than two genes (usually by many different genes) at different loci on different chromosomes. These genes are described as polygenes. Examples of human polygenic inheritance are height, skin colour and weight. Polygenes allow a wide range of physical traits. For instance, height is regulated by several genes so that there will be a wide range of heights in a population.
U10.2.5 Chi-squared tests are used to determine whether the difference between an observed and expected frequency distribution is statistically significant.
UNDERSTANDINGS:
U10.2.1 Gene loci are said to be linked if on the same chromosome.
Linkage refers to genes that are located on the same chromosome. Linked genes tend to be inherited together and the extent of crossing over depends on how close together they are on the chromosome. Linkage generally reduces the variety of offspring that can be produced. In genetic crosses, linkage is indicated when a greater proportion of the offspring resulting from a cross are of the parental type (than would be expected if the alleles were assorting independently). If the genes in question had been on separate chromosomes, there would have been more genetic variation in the gametes and therefore in the offspring.
U10.2.2 Unlinked genes segregate independently as a result of meiosis.
- Unlinked genes are on different chromosomes
- At Metaphase 1, each pair of homologous chromosomes line up at the equator of the spherical cell
- For each homologous pair, the chromosomes assort to one side or the other
- The alignment of any pair of homologous chromosomes is independent of all the other pairs
- Thus, the mixture of unlinked genes in gametes is random.
U10.2.3 Variation can be discrete or continuous.
Discontinuous variation This is where individuals fall into a number of distinct classes or categories, and is based on features that cannot be measured across a complete range. You either have the characteristic or you don't. Blood groups are a good example: you are either one blood group or another - you can't be in between, same goes for gender. Such data is called discrete (or categorical) data. Chi-squared statistical calculations work well in this case.
In continuous variation there is a complete range of measurements from one extreme to the other. Height is an example of continuous variation - individuals can have a complete range of heights, for example, 1.6, 1.61, 1.62, 1.625 etc metres high.
U10.2.4 The phenotypes of polygenic characteristics tend to show continuous variation.
A10.2.3 Polygenic traits such as human height may also be influenced by environmental factors.
Continuous variation is the combined effect of many genes (known as polygenic inheritance) and is often significantly affected by environmental influences. Milk yield in cows, for example, is determined not only by their genetic make-up but is also significantly affected by environmental factors such as pasture quality and diet, weather, and the comfort of their surroundings.
Polygenic traits are controlled by two or more than two genes (usually by many different genes) at different loci on different chromosomes. These genes are described as polygenes. Examples of human polygenic inheritance are height, skin colour and weight. Polygenes allow a wide range of physical traits. For instance, height is regulated by several genes so that there will be a wide range of heights in a population.
U10.2.5 Chi-squared tests are used to determine whether the difference between an observed and expected frequency distribution is statistically significant.
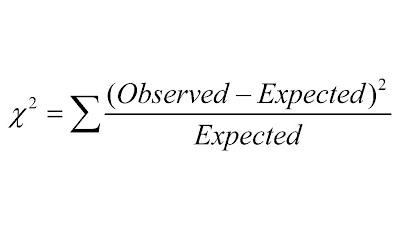
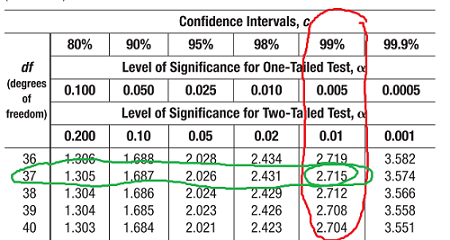
Degree of freedom - the number of independent values or quantities that can be
assigned to a statistical distribution.
To find the degree of freedom = n - 1, where n is the number of data points
A critical value is the number you compare your own result to (critical values would be provided in a test).
A confidence interval does not quantify variability. A 95% confidence interval is a range of values that you can be 95% certain contains the true mean of the population.
(Ignore the circled row and column)
If your OWN value for something is greater than the critical value, you REJECT the null hypothesis and ACCEPT the experimental hypothesis.
If you OWN value for something is less than the critical value, you ACCEPT the null hypothesis and REJECT the experimental hypothesis.
APPLICATION:
A10.2.1 Morgan’s discovery of non-Mendelian ratios in Drosophila.
1. Male fly with white eyes (flies usually have red eyes) was crossed with a female fly with red eyes (a wild type)
2. The offspring of the first generation were red eyed
3. Males and females of the first generation were crossed with each other
4. The white-eye trait reappeared in the expected 3:1 Mendelian ratio for a recessive trait. However only the males had white eyes
5. This suggested that the white-eye trait is carried on the X chromosome
6. Crossing white-eyed males and red-eyed females from the second generation produced equal numbers of offspring with each eye colour
7. Males have white eyes when they inherit the mutant gene on the X chromosome from their mother. Females only show the trait if they inherit mutant genes on both X chromosomes. This became known as "sex linkage"
A10.2.2 Completion and analysis of Punnett squares for dihybrid traits.
S10.2.1 Calculation of the predicted genotypic and phenotypic ratio of offspring of dihybrid crosses involving unlinked autosomal genes.
Dihybrid Cross Example
Next, write out the possible gametes: SB, SB, sb and sb or simply just SB and sb
To find the degree of freedom = n - 1, where n is the number of data points
A critical value is the number you compare your own result to (critical values would be provided in a test).
A confidence interval does not quantify variability. A 95% confidence interval is a range of values that you can be 95% certain contains the true mean of the population.
(Ignore the circled row and column)
If your OWN value for something is greater than the critical value, you REJECT the null hypothesis and ACCEPT the experimental hypothesis.
If you OWN value for something is less than the critical value, you ACCEPT the null hypothesis and REJECT the experimental hypothesis.
APPLICATION:
A10.2.1 Morgan’s discovery of non-Mendelian ratios in Drosophila.
1. Male fly with white eyes (flies usually have red eyes) was crossed with a female fly with red eyes (a wild type)
2. The offspring of the first generation were red eyed
3. Males and females of the first generation were crossed with each other
4. The white-eye trait reappeared in the expected 3:1 Mendelian ratio for a recessive trait. However only the males had white eyes
5. This suggested that the white-eye trait is carried on the X chromosome
6. Crossing white-eyed males and red-eyed females from the second generation produced equal numbers of offspring with each eye colour
7. Males have white eyes when they inherit the mutant gene on the X chromosome from their mother. Females only show the trait if they inherit mutant genes on both X chromosomes. This became known as "sex linkage"
A10.2.2 Completion and analysis of Punnett squares for dihybrid traits.
S10.2.1 Calculation of the predicted genotypic and phenotypic ratio of offspring of dihybrid crosses involving unlinked autosomal genes.
- A dihybrid cross is a cross between two individuals that shows the inheritance of two different genes at the same time; usually involving unlinked autosomal genes.
- Note: The following example contains two unlinked genes, which means the genes are on different chromosomes. This means they follow Mendel’s law of independent assortment.
Dihybrid Cross Example
- Since almost all examples look at Mendel’s pea plants, for this example we will look at two traits in cats; hair length and color.
- If the question stated that short hair (S) is dominant over long hair (s) and black fur (B) is dominant over white fur (b), what would be the genotypic and phenotypic ratio of a cross between a short hair black cat that is homologous for these traits and a long hair white cat?
Next, write out the possible gametes: SB, SB, sb and sb or simply just SB and sb
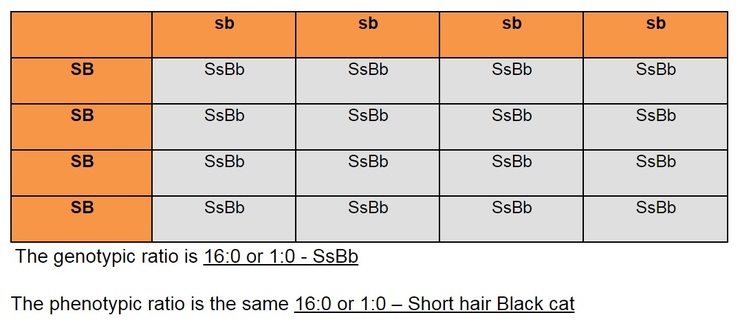
S10.2.2 Identification of recombinants in crosses involving two linked genes.
Linked genes that have undergone recombination can be distinguished from unlinked genes via a test cross because the frequency of the recombinant genotypes will always be less than would occur for unlinked genes (crossing over does not happen every time)
For example:
S10.2.3 Use of a chi-squared test on data from dihybrid crosses.
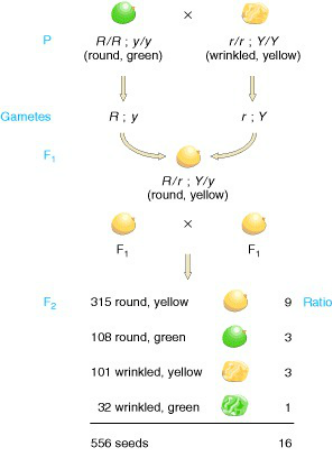
For an example, use Mendel’s results from his pea plant crosses.
When he did a dihybrid cross between two heterozygotes RrYy x RrYy, the expected phenotypic ratio due to independent assortment would be 9:3:3:1. Look at the chart below to see his actual results.
The Ho (null hypothesis) would be the results are due to independent assortment and the Ha (alternative hypothesis) would be that the alleles do not assort independently and the results are due to gene linkage.
Linked genes that have undergone recombination can be distinguished from unlinked genes via a test cross because the frequency of the recombinant genotypes will always be less than would occur for unlinked genes (crossing over does not happen every time)
For example:
- Heterozygous test cross of unlinked genes = 1 : 1 : 1 : 1 phenotypic ratio
- Heterozygous test cross of linked genes = 1 : 1 : 0.1 : 01 phenotypic ratio (uncommon phenotypes are recombinants)
S10.2.3 Use of a chi-squared test on data from dihybrid crosses.
For an example, use Mendel’s results from his pea plant crosses.
When he did a dihybrid cross between two heterozygotes RrYy x RrYy, the expected phenotypic ratio due to independent assortment would be 9:3:3:1. Look at the chart below to see his actual results.
The Ho (null hypothesis) would be the results are due to independent assortment and the Ha (alternative hypothesis) would be that the alleles do not assort independently and the results are due to gene linkage.
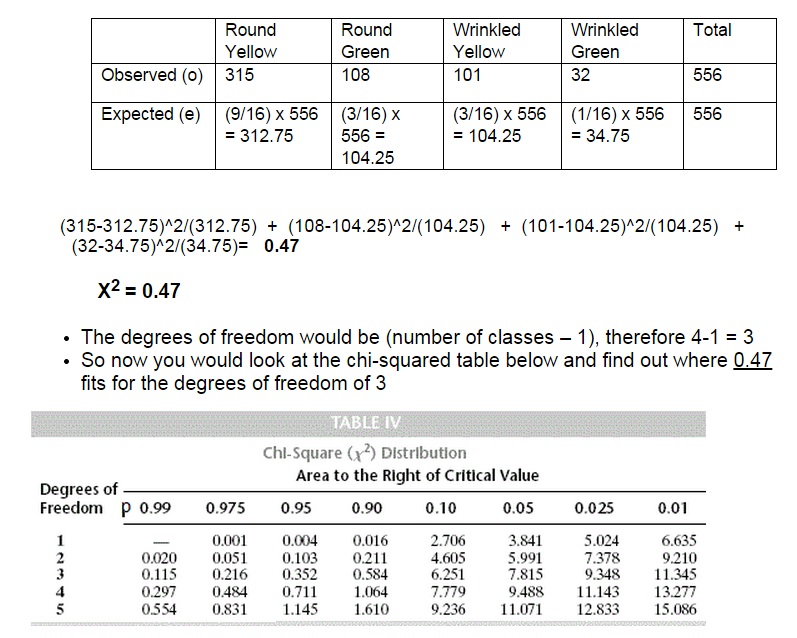
As you can see from the table above the critical value at the 0.05 level of significance is 7.815.
What the p-value of 0.05 or 5% indicates is the probability of getting the results you did (or more extreme results) given that the null hypothesis is true.
If the calculated value is above or equal to 7.815 then we reject the Ho and accept the HA that the alleles in question are linked.
However, since our chi-squared value of 0.47 is significantly less than this value (it has a p-value of 0.90 or 90%) we accept the Ho that the results that Mendel saw were due to independently assortment of the alleles.
What the p-value of 0.05 or 5% indicates is the probability of getting the results you did (or more extreme results) given that the null hypothesis is true.
If the calculated value is above or equal to 7.815 then we reject the Ho and accept the HA that the alleles in question are linked.
However, since our chi-squared value of 0.47 is significantly less than this value (it has a p-value of 0.90 or 90%) we accept the Ho that the results that Mendel saw were due to independently assortment of the alleles.
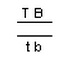
NOTE: Alleles
are usually shown side by side in dihybrid crosses, for example, TtBb. In
representing crosses involving linkage, it is more common to show them as
vertical pairs, for example:
Essential idea: Gene pools change over time.
10.3 Gene pools and speciation
UNDERSTANDINGS:
U10.3.1 A gene pool consists of all the genes and their different alleles, present in an interbreeding population.
Allele frequency: the proportion of an allele within a population
Gene pool: the sum total of alleles present in a sexually reproducing population.
U10.3.2 Evolution requires that allele frequencies change with time in populations.
Evolution is the cumulative change in allele frequency or heritable characteristics in a population over time. The cumulative change can occur as a result of genetic mutations and selective pressures which favour certain heritable characteristics over other less favourable characteristics.
These populations have to be reproductively isolated, thus preventing gene flow between populations. If a population that has a certain allele or characteristic is quite small, random events such as disease or natural disasters can cause a drastic drop in this particular allele.
U10.3.3 Reproductive isolation of populations can be temporal, behavioural or geographic.
PRE-ZYGOTIC BARRIERS (barriers that prevent mating or fertilisation):
Temporal isolation - occurs when two species mate or flower at different times of the year.
Behavioural isolation - occurs when two species respond to different specific courtship patterns.
Geographic isolation - occurs when two populations of the same species or breeding group are separated by a physical barrier, such as a mountain or body of water.
Ecological isolation - occurs when two species inhabit similar regions, but occupy different habitats.
Mechanical isolation - occurs when genital differences prevent copulation (animals) or when flowers are pollinated by different animals (plants).
POST-ZYGOTIC BARRIERS (barriers that prevent hybrid zygote turning into a strong adult):
Hybrid inviability - hybrids are produced but fail to develop to reproductive maturity.
Hybrid infertility - hybrids fail to produce functional gametes.
Hybrid breakdown - the F1 (first generation) hybrids are fertile but the F2 fail to develop or are infertile.
U10.3.4 Speciation due to divergence of isolated populations can be gradual.
Divergent evolution:
U10.3.5 Speciation can occur abruptly.
Punctuated equilibrium implies long periods without appreciable change and short periods of rapid evolution:
A10.3.1 Identifying examples of directional, stabilizing and disruptive selection.
DIRECTIONAL:
In directional selection, one extreme of the trait distribution experiences selection against it. The result is that the population's trait distribution shifts toward the other extreme. In the case of such selection, the mean of the population graph shifts. Using the familiar example of giraffe necks, there was a selection pressure against short necks, since individuals with short necks could not reach as many leaves on which to feed. As a result, the distribution of neck length shifted to favor individuals with long necks.
UNDERSTANDINGS:
U10.3.1 A gene pool consists of all the genes and their different alleles, present in an interbreeding population.
Allele frequency: the proportion of an allele within a population
Gene pool: the sum total of alleles present in a sexually reproducing population.
U10.3.2 Evolution requires that allele frequencies change with time in populations.
Evolution is the cumulative change in allele frequency or heritable characteristics in a population over time. The cumulative change can occur as a result of genetic mutations and selective pressures which favour certain heritable characteristics over other less favourable characteristics.
These populations have to be reproductively isolated, thus preventing gene flow between populations. If a population that has a certain allele or characteristic is quite small, random events such as disease or natural disasters can cause a drastic drop in this particular allele.
U10.3.3 Reproductive isolation of populations can be temporal, behavioural or geographic.
PRE-ZYGOTIC BARRIERS (barriers that prevent mating or fertilisation):
Temporal isolation - occurs when two species mate or flower at different times of the year.
Behavioural isolation - occurs when two species respond to different specific courtship patterns.
Geographic isolation - occurs when two populations of the same species or breeding group are separated by a physical barrier, such as a mountain or body of water.
Ecological isolation - occurs when two species inhabit similar regions, but occupy different habitats.
Mechanical isolation - occurs when genital differences prevent copulation (animals) or when flowers are pollinated by different animals (plants).
POST-ZYGOTIC BARRIERS (barriers that prevent hybrid zygote turning into a strong adult):
Hybrid inviability - hybrids are produced but fail to develop to reproductive maturity.
Hybrid infertility - hybrids fail to produce functional gametes.
Hybrid breakdown - the F1 (first generation) hybrids are fertile but the F2 fail to develop or are infertile.
U10.3.4 Speciation due to divergence of isolated populations can be gradual.
Divergent evolution:
- Species descended from a common ancestor gradually diverge more and more in morphology i.e. become phenotypically different - adaptive radiation, through the slow but relentless effects of natural selection.
U10.3.5 Speciation can occur abruptly.
Punctuated equilibrium implies long periods without appreciable change and short periods of rapid evolution:
- A new species changes most when it first diverges from a parent species and then changes little for the rest of its existence.
A10.3.1 Identifying examples of directional, stabilizing and disruptive selection.
DIRECTIONAL:
In directional selection, one extreme of the trait distribution experiences selection against it. The result is that the population's trait distribution shifts toward the other extreme. In the case of such selection, the mean of the population graph shifts. Using the familiar example of giraffe necks, there was a selection pressure against short necks, since individuals with short necks could not reach as many leaves on which to feed. As a result, the distribution of neck length shifted to favor individuals with long necks.
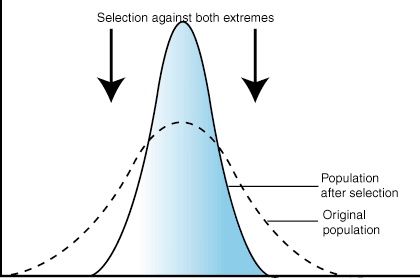
STABILIZING:
When selective pressures select against the two extremes of a trait, the population experiences stabilizing selection. For example, plant height might be acted on by stabilizing selection. A plant that is too short may not be able to compete with other plants for sunlight. However, extremely tall plants may be more susceptible to wind damage. Combined, these two selection pressures select to maintain plants of medium height. The number of plants of medium height will increase while the numbers of short and tall plants will decrease.
When selective pressures select against the two extremes of a trait, the population experiences stabilizing selection. For example, plant height might be acted on by stabilizing selection. A plant that is too short may not be able to compete with other plants for sunlight. However, extremely tall plants may be more susceptible to wind damage. Combined, these two selection pressures select to maintain plants of medium height. The number of plants of medium height will increase while the numbers of short and tall plants will decrease.
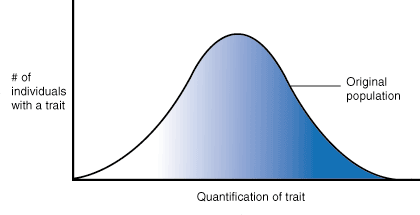
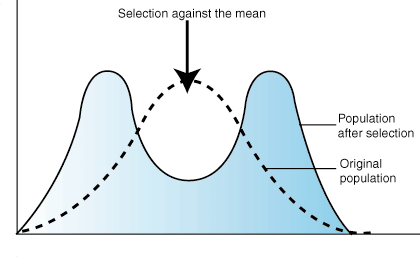
DISRUPTIVE:
In disruptive selection, selection pressures act against individuals in the middle of the trait distribution. The result is a bimodal, or two-peaked, curve in which the two extremes of the curve create their own smaller curves. For example, imagine a plant of extremely variable height that is pollinated by three different pollinators, one that was attracted to short plants, another that preferred plants of medium height and a third that visited only the tallest plants. If the pollinator that preferred plants of medium height disappeared from an area, medium height plants would be selected against and the population would tend toward both short and tall, but not medium height plants. Such a population, in which multiple distinct forms or morphs exist is said to be polymorphic.
In disruptive selection, selection pressures act against individuals in the middle of the trait distribution. The result is a bimodal, or two-peaked, curve in which the two extremes of the curve create their own smaller curves. For example, imagine a plant of extremely variable height that is pollinated by three different pollinators, one that was attracted to short plants, another that preferred plants of medium height and a third that visited only the tallest plants. If the pollinator that preferred plants of medium height disappeared from an area, medium height plants would be selected against and the population would tend toward both short and tall, but not medium height plants. Such a population, in which multiple distinct forms or morphs exist is said to be polymorphic.
A10.3.2 Speciation in the genus Allium by polyploidy.
Speciation - the formation of new and distinct species in the course of evolution.
Polyploidy is a condition in which an organism has more than two complete sets of chromosomes in all somatic cells i.e. > diploid. It is far more common in plant species as they lack separate sexes and are capable of asexual reproduction (self-pollination). It may occur as a result of the failutre of a meiotic cell to undergo cytpkinesis (so chromosome replication occurs minus cell division). Consequently, gametes are diploid (2n) and resulting offspring are tetraploid (4n). Because tetraploid offspring can no longer mate with diploid organisms (triploid offspring tend to be infertile), speciation has occured.
S10.3.1 Comparison of allele frequencies of geographically isolated populations.
-
Speciation - the formation of new and distinct species in the course of evolution.
Polyploidy is a condition in which an organism has more than two complete sets of chromosomes in all somatic cells i.e. > diploid. It is far more common in plant species as they lack separate sexes and are capable of asexual reproduction (self-pollination). It may occur as a result of the failutre of a meiotic cell to undergo cytpkinesis (so chromosome replication occurs minus cell division). Consequently, gametes are diploid (2n) and resulting offspring are tetraploid (4n). Because tetraploid offspring can no longer mate with diploid organisms (triploid offspring tend to be infertile), speciation has occured.
S10.3.1 Comparison of allele frequencies of geographically isolated populations.
-
ADDITIONAL DEFINITIONS AND DETAILS:
- Hybrid is an offspring resulting from the cross between parents of different species or sub-species.
- Polyploidy can lead to increased size, resistance to disease and overall vigour (advantageous to plants with polyploidy).
- Autopolyploidy -> polyploids with multiple chromosomes derived from a single species - from the fusion of 2n gametes (asexual reproduction).
- Allopolyploidy -> an individual whose chromosomes are composed of more than two genomes, each of which has been derived complete but possibly modified from one of the two or more species.
- Allopolyploids come about when a sterile F1 (first generation) hybrid doubles all of its chromosomes and become fertile.
- Allopatric speciation -> allo = different; patric = fatherland - arises when a species is subject too geographic isolation and can occur when a population is split by a physical barrier such as:
- Sympatric speciation ->sym = same; patric = fatherland - the formation of two or more descendant species from a single ancestral species all occupying the same geographic location.
- Convergent evolution -> describes the aquisition of the same biological trait in unrelated lineages. They evolved independently in organisms only very distant related, leading to analogous structures.
- Divergent evolution -> adaptive radiation - derived from a common ancestor and become phenotypically different, leading to homologous structures.
- Hybrid is an offspring resulting from the cross between parents of different species or sub-species.
- Polyploidy can lead to increased size, resistance to disease and overall vigour (advantageous to plants with polyploidy).
- Autopolyploidy -> polyploids with multiple chromosomes derived from a single species - from the fusion of 2n gametes (asexual reproduction).
- Allopolyploidy -> an individual whose chromosomes are composed of more than two genomes, each of which has been derived complete but possibly modified from one of the two or more species.
- Allopolyploids come about when a sterile F1 (first generation) hybrid doubles all of its chromosomes and become fertile.
- Allopatric speciation -> allo = different; patric = fatherland - arises when a species is subject too geographic isolation and can occur when a population is split by a physical barrier such as:
- a river
- a mountain range
- a desert
- a road
- the sea etc.
- Sympatric speciation ->sym = same; patric = fatherland - the formation of two or more descendant species from a single ancestral species all occupying the same geographic location.
- Convergent evolution -> describes the aquisition of the same biological trait in unrelated lineages. They evolved independently in organisms only very distant related, leading to analogous structures.
- Divergent evolution -> adaptive radiation - derived from a common ancestor and become phenotypically different, leading to homologous structures.